library(tidyverse)
library(glmmTMB)
library(easystats)
intell <- read_csv("intell_sim.csv")
glimpse(intell)Rows: 1,800
Columns: 5
$ subject <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2…
$ icontrol <dbl> -0.0848893, -0.0848893, -0.0848893, -0.0848893, -0.0848893…
$ trial <dbl> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 1, 2, 3, 4, 5, 6, 7…
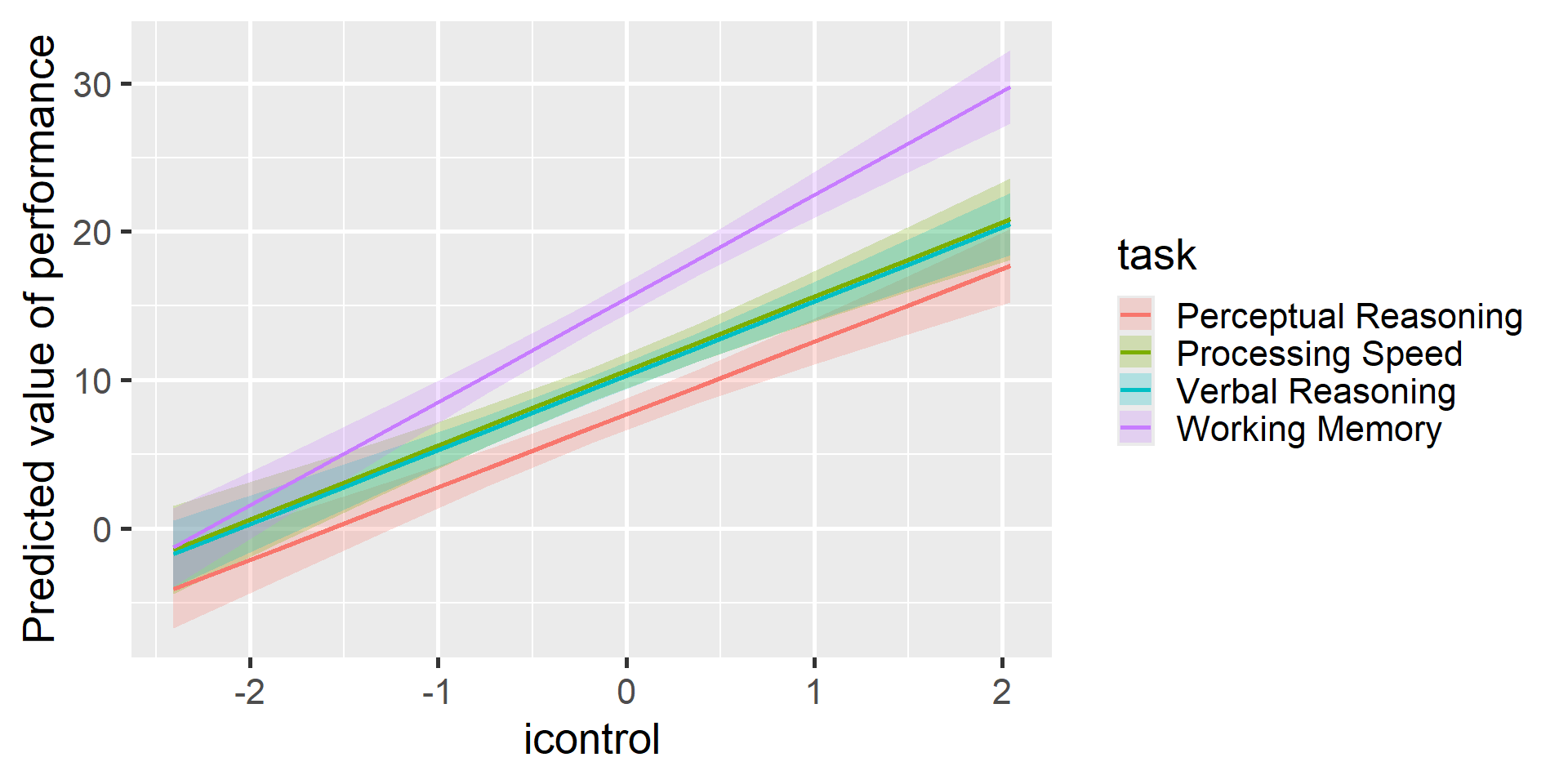
$ task <chr> "Perceptual Reasoning", "Processing Speed", "Working Memor…
$ performance <dbl> 17.997978, 14.412448, 22.439953, 17.173733, 21.686384, 14.…